Use this to specify options for a compatibility check of QTL data.
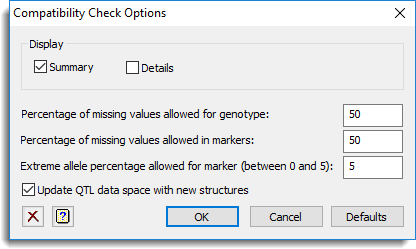
Display
This specifies which items of output are to be produced by the analysis.
| Summary | A summary of the changes |
| Details | Displays details of the replaced structures |
Percentage of missing values allowed for genotypes
This specifies a threshold on the percentage of missing values within each genotype. Genotypes with more than that percentage of missing marker scores are excluded. The genotypes that are excluded will also be excluded from the corresponding values in other structures such as the traits, the genotypes factor, the kinship matrix and the subpopulations factor when specified on the main menu.
Percentage of missing values allowed for in markers
This specifies a threshold on the percentage of missing values allowed within each marker. Markers with more than that percentage of missing scores will be excluded.
Extreme allele percentage allowed for marker
This specifies that markers will be excluded if one allele percentage of that marker is less than the this value.
Update QTL data space with new structures
When selected the new structures will be updated or added to the QTL data space.
Action buttons
| OK | Stores the option settings and closes the dialog. |
| Cancel | Close the dialog without making any changes. |
| Defaults | Sets the options to their default settings. |
Action Icons
| Clear | Clear all fields and list boxes. | |
| Help | Open the Help topic for this dialog. |
See also
- Compatibility check menu.
- QTL data space for using data in QTL menus
- QMATCH procedure in command mode