Select menu: Run | Submit BUGS Script
WinBUGS (Bayesian inference Using Gibbs Sampling, Spiegelhalter, Thomas, Best & Lund 2003) is an application that can be used for the Bayesian analysis of complex models using Markov Chain Monte Carlo (MCMC) methods.
WinBUGS is available free at https://www.mrc-bsu.cam.ac.uk/software/bugs/the-bugs-project-winbugs/ and an open-source version of the core BUGS code (OpenBUGS) is also available at http://www.openbugs.net/. Use this to run WinBUGS or OpenBUGS from Genstat in batch mode using scripts. To execute commands within WinBUGS, Genstat automatically creates a data and script file containing the necessary commands for the MCMC. These files are passed to WinBUGS and, once it has completed execution, the results can be imported into Genstat.
- From the menu select Run | Submit BUGS Script.
- You can set additional Options before running the analysis.
- Enter values by double-clicking them in the Available data list or select and move them across using
 then click Run.
then click Run.
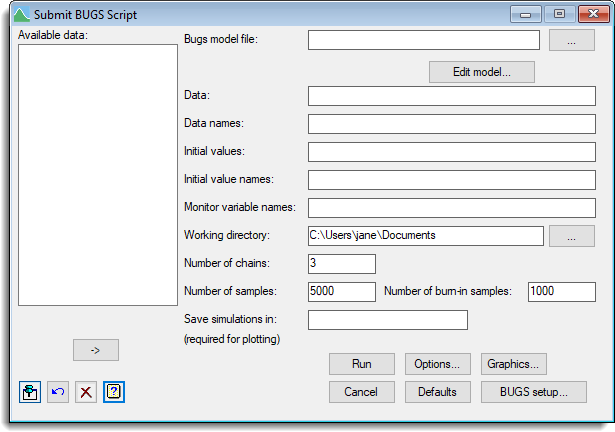
To use this menu either WinBUGS or OpenBUGS must be installed on the current system. The control of whether WinBUGS or OpenBUGS is to be run and the location of the executable should be supplied within the BUGS setup dialog. WinBUGS requires the script file generated by Genstat to be written to the directory containing the executable. Therefore, you must have write permission to this directory to be able to run.
BUGS model file
Specifies a filename containing the model to run in WinBUGS or OpenBUGS. The model must be supplied in a text file using BUGS code. Details of how to write models can be found in the online help within WinBUGS or OpenBUGS. You can browse for a filename by clicking on the ![]() button.
button.
Edit model
This opens the Edit BUGS Model dialog which allows you to edit the text defining the BUGS model.
Data
Specifies the data for the analysis. The data can either be supplied as a pointer where each element of the pointer represents a different data structure used in the model or by supplying the names of the different data structures. The data structures can be a scalar or a variate or, for 2-D data, a matrix or pointer to variates.
Data names
A text or list supplying the names for the data structures. The names should be in the same order as that in which the data structures occur in the data.
Initial values
Specifies a list of pointers containing the initial values for each of the chains. The elements of each pointer can be either a scalar or variate, and must be in the same order in all the pointers.
Initial value names
A list of the names of the variables for the initial values where the names are in the same order as the data within the pointers for the initial values. Alternatively, a text can be supplied containing all the names.
Monitor variable names
A list of the names of the variables of interest that are to be monitored. Alternatively, a text can be supplied containing all the names.
Working directory
The working directory to be used.
Number of chains
The number of Markov chains to be run.
Number of samples
The number of samples to run after the burn-in.
Number of samples for burn-in
Length of burn-in for each chain.
Save simulations in
Specify an identifier name to save the simulations. The results are saved in a list of pointers, where each pointer contains the simulations and associated information for a Markov chain. If further analysis is required to view plots or diagnostics then the simulation results must be saved using this option.
Action buttons
| Run | Run the analysis. |
| Cancel | Close the menu without further changes. |
| Options | Opens a dialog where additional options and settings can be specified for the analysis. |
| Defaults | Set the menu settings back to the default settings. Clicking the right mouse on this button produces a shortcut menu where you can choose to set the options using the currently stored defaults or the Genstat default settings. |
| Graphics | Opens a dialog where you can plot results and diagnostics from the MCMC analysis. |
Action Icons
| Pin | Controls whether to keep the dialog open when you click Run. When the pin is down |
|
| Restore | Restore names into edit fields and default settings. | |
| Clear | Clear all fields and list boxes. | |
| Help | Open the Help topic for this dialog. |
See also
- Edit BUGS Model menu.
- Submit BUGS Script Options menu.
- BUGS Setup menu.
- BUGS plots menu.
- BGXGENSTAT procedure.