Use this to partition a genotype factor into test genotypes (i.e. genotypes that are to be taken forward to QTL analysis) and extra genotypes present in the trial (eg. check lines and/or parents). This dialog appears when you select a genotype to partition and click OK.
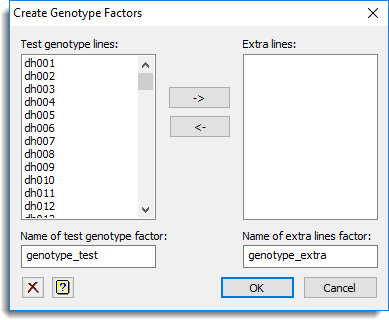
The dialog generates two factors, identifying the test genotypes and extra lines, which can then be used in the preliminary single environment analysis menu. The generated test genotype factor has levels/labels corresponding to the genotypes to be used in the QTL analysis, with missing values used to represent the extra lines. The generated extra line factor has levels/labels corresponding to the extra lines plus an additional level/label (e.g. level 0 and/or label ‘Test line’) used to identify the set of genotypes taken forward for QTL analysis. For example, if a genotype factor contains 22 genotypes, where levels 21 and 22 represent the two parents, the two factors will be generated as follows:
| Test genotype factor | Extra lines factor |
| 1 | 0 |
| 2 | 0 |
| 3 | 0 |
| … | … |
| 20 | 0 |
| * | 21 |
| * | 22 |
Test genotype lines
This lists the levels or labels of the genotype factor that represent the test genotypes. Levels or labels can be copied across to the Extra lines list by double-clicking on an item or by making a selection and clicking the ![]() button.
button.
Name of test genotype factor
Specifies an identifier name to use for the test genotype factor. The generated test genotype factor will insert missing values in units corresponding to check lines.
Extra lines
This lists the levels or labels of the genotype factor that represent the extra lines. Levels or labels can be copied across to the Test genotype lines list by double-clicking on an item or by making a selection and clicking the ![]() button.
button.
Name of extra lines factor
Specifies an identifier name to use for the extra lines factor. The generated check line factor will create a new level (0) and/or label (Test line) to represent the test genotypes.
Action Icons
| Clear | Clear all fields and list boxes. | |
| Help | Open the Help topic for this dialog. |
See also
- Preliminary single environment analysis menu for analysis of experiments for QTL analysis
- QTL data space for using data in QTL menus