Use this to save results associated with the test line genotypes from a Preliminary single environment analyses. The data saved using this dialog can be used for genotype by environment and QTL analysis. For a multi-environment analysis use this to stack the results from each individual trial analysis where each set of results is identified by the environment label.
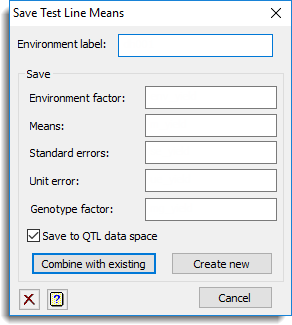
Environment label
Specifies a label that can be used to identify the results from an analysis for a particular environment. The label is used when creating or appending a new level to the Environment factor. Note that this label is required even if the analysis is for a single environment trial.
Save
Specifies the different data structures that can be saved after each analysis.
| Environment factor | Factor | Creates a factor where each level represents the results from a different environment. The factor is created using the label supplied in the Environment label field. |
| Means | Variate | Saves the predicted means for the test lines. |
| Standard errors | Variate | Saves the standard errors of the predicted genotype means. |
| Unit error | Variate | Saves the unit error to represent uncertainty on trait means (derived from individual unit or plot error) to be included in a QTL analysis. |
| Genotypes | Factor | Saves the test line genotypes. |
Save to QTL data space
When the results are saved the data structure names will be added to the QTL data space so that the names of the structures will be present in the QTL menus.
Combine with existing
This can be used to stack results from different analysis together. When results from trials are stacked together the environment factor identifies the results from each trial. The data are stacked and expanded to include a full set of results based on the genotypes that are present in the stacked environments. To stack results the names used to save the data must be the same for each analysis whose results are to be combined.
Create new
This will create a new set of results overwriting any existing data structures.
Action Icons
| Clear | Clear all fields and list boxes. | |
| Help | Open the Help topic for this dialog. |
See also
- Preliminary single environment analysis menu for analysis of experiments for QTL analysis
- QTL data space for using data in QTL menus
- VPREDICT for predicting means from a mixed model analysis