Use this to save results from an automatic analysis of series of trials in Genstat data structures. The individual trial or the meta analysis of all trials must be selected using the Model to save results for dropdown list.
- After selecting the appropriate boxes, type the identifiers of the data structures into the corresponding In: fields.
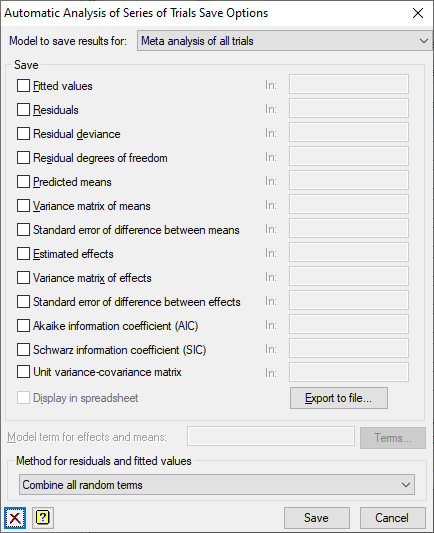
Model to save results for
The results are saved for the selected trial or the meta analysis. The meta analysis option will be available only if the Run a meta analysis to combine information across trials option has been chosen in the Options dialog,
Save
| Fitted values | Variate | Fitted values from the analysis |
| Residuals | Variate | Residuals from the analysis |
| Residual deviance | Scalar | Residual deviance from fitting the full fixed model |
| Residual degrees of freedom | Scalar | Residual degrees of freedom after fitting the full fixed model |
| Predicted means | Table | Predicted means for the specified term |
| Variance matrix of means | Symmetric matrix | Variance-covariance matrix for the predicted means |
| Standard error of difference between means | Symmetric matrix | Standard errors of differences between the predicted means |
| Estimated effects | Table | Estimated regression coefficients for the specified term |
| Variance matrix of effects | Symmetric matrix | Variance-covariance matrix for the estimated effects |
| Standard error of difference between effects | Symmetric matrix | Standard errors of differences between the estimated parameters of each term |
| Akaike information coefficient (AIC) | Scalar | Akaike information coefficient to assess the random model |
| Schwarz information coefficient (SIC) | Scalar | Schwarz information coefficient to assess the random model |
| Unit variance-covariance matrix | Symmetric matrix | Matrix giving the fitted variance-covariance relationship between all the units. Note: this may be very large. This uses the random and residual terms, but not spline terms. It cannot be formed if the model contains sparse inverse covariance matrices. |
Display in spreadsheet
Select this to display the results in a new spreadsheet window.
Export to file
Save selected results into a Genstat or Excel file using the Save REML results in a spreadsheet dialog
Model terms for effects and means
Specifies the model term for which tables of estimated effects and predicted means are to be saved. The string ‘Constant’ may be used to save results for the constant term. If tables of effects or means are required for more than one model term, this menu should be invoked once for each term, changing the specification of the model term each time.
Method for residuals
The list allows selection from type of residuals that can be calculated.
| Combine all random terms | Use the residuals combined from all random terms. |
| Final random term only | Use the residuals from the final random term. |
| Standardized residuals from all random terms | Uses standardized residuals after combining them from all random terms. |
| Standardized residuals from final random term only | Uses standardized residuals from the final random term. |
| Combine all random terms, excluding spline terms | Use the residuals combined from all random terms except spline terms. |
See also
- Save REML results in a spreadsheet
- Automatic Analysis of Series of Trials menu
- Options for specifying output options
- Further Output for obtaining additional output
after fitting a model - Residual Plots for generating plots of residuals
- Means Plots for generating plots of one- or two-way tables of means
- REML directive for command mode use of REML, with additional options to
control the algorithm and for more sophisticated analyses - VCOMPONENTS directive for further information about
fixed, random, and spline model terms - Linear Mixed Models (REML) – Correlated Errors for setting up covariance models
- Reml Predictions menu for forming predictions
- VPLOT procedure for plotting residuals
- VFRESIDUALS procedure to calculate standardized residuals