Select menu: Stats | Multivariate Analysis | Principal Coordinates
- After you have imported your data, from the menu select
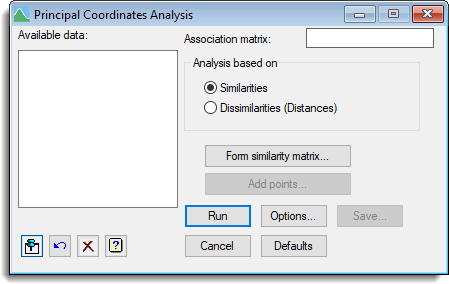
Stats | Multivariate Analysis | Principal Coordinates. - Fill in the fields as required then click Run.
You can set additional Options then after running, you can save the results by clicking Save.
Given a symmetric matrix, A say, with values representing the associations amongst a set of n units, principal coordinates analysis attempts to find a set of points for the n units in a multidimensional space so that the squared distance between the ith and jth points is given by:
dij = aii + ajj – 2aij
If, for example, A is a similarity matrix then aii and ajj are both equal to 1.0, and so this is equivalent to:
dij = 2 (1 – aij)
Thus similar units are placed close together and dissimilar units are further apart.
The coordinates generated can be arbitrarily located in space as this will not alter the fitted inter-point distances. By convention, the points are centred to have their mean at the origin, and rotated to principal axes, so that the first r dimensions give the best r dimensional fit.
Association matrix
This specifies the name of the symmetric matrix of associations to be analysed and can be selected from Available data. You can specify either a similarity or distance matrix.
Form similarity matrix
This forms a similarity matrix for input to principal coordinates. You should select this menu if you are starting with raw data variates. This opens the Form Similarity Matrix dialog.
Analysis based on
Specifies whether the analysis is based on a similarity matrix or distance matrix. Whereas similarity matrices can be input directly to the analysis, a dissimilarity (distance) matrix requires preliminary transformation. Selecting dissimilarities will ensure the appropriate transformation is applied to the data.
If B is a distance matrix, so that bij gives the observed distance between the ith and jth units, then the transformation
A = – B * B / 2
will lead to the analysis generating points with inter-point squared distance
dij = aii + ajj – 2aij = 0 + 0 – 2 * (- bij * bij / 2 ) = bij2
Therefore the analysis will give points that generate the supplied distances; the first r dimensions of the solution will give the best r-dimensional fit.
Action buttons
| Run | Run the analysis. |
| Cancel | Close the dialog without further changes. |
| Options | Opens a dialog where additional options and settings can be specified for the analysis. |
| Defaults | Reset options to the default settings. Clicking the right mouse on this button produces a pop-up menu where you can choose to set the options using the currently stored defaults or the Genstat default settings. |
| Save | Opens a dialog to specify names of structures to save the results from the analysis. |
Action Icons
| Pin | Controls whether to keep the dialog open when you click Run. When the pin is down |
|
| Restore | Restore names into edit fields and default settings. | |
| Clear | Clear all fields and list boxes. | |
| Help | Open the Help topic for this dialog. |
See also
- Options for choosing which results to display.
- Saving results for further analysis.
- Form Similarity Matrix menu.
- Adding new points to a principal coordinates analysis.
- PCO for additional data options in command mode.
- ADDPOINTS directive for adding new points to a principal coordinates analysis in command mode.
- Multidimensional scaling for an alternative method of fitting distances.