This can be used to add or remove marker genotype scores and associated data structures in the QTL data space.
- To view this dialog click
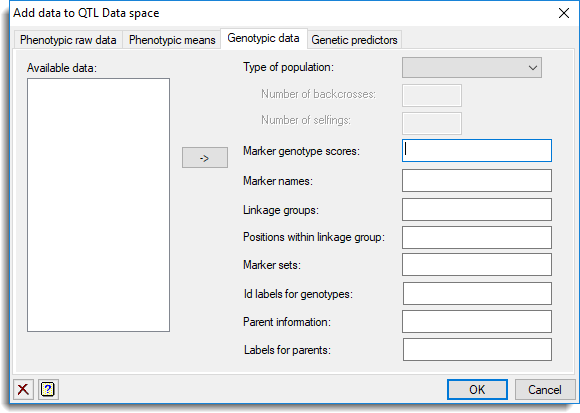
 within the QTL Data view then click the Genetic data tab.
within the QTL Data view then click the Genetic data tab.

If data structure names have been stored in a QTL data space then when the QTL menus are opened the data structure names will automatically be entered into the relevant fields. Also only the data structures present within that data space will be displayed in the Available data, otherwise all the current data within Genstat will be displayed. When data are present within the QTL data space you can right-click on the Available data list to open a shortcut menu where you can change between displaying data only within the data space and all data within Genstat.
Available data
This lists data structures currently available in Genstat appropriate to the current input field. Double-click a name or by making a selection and clicking the ![]() button to copy the name to the current input field or type the name.
button to copy the name to the current input field or type the name.
Type of population
A list of population types. Select populations as follows:
- F2 for an F2 population
- BC1 for a backcross population
- DH for a double-haploid population
- RILn for a population of recombinant inbred lines
- BCxSy for a population of backcross inbred lines
- CP for cross pollinator
- AMP for an association mapping population.
If RILn is selected the number of generations for the population (n) can be supplied using the Number of generations option. If BCxSy is selected the number of backcrosses and selfings for the population can be supplied using the Number of backcrosses and Number of selfings options respectively. When a population is selected the relevant fields to supply data structures will be enabled on the dialog.
Marker genotype scores
A pointer specifying the marker scores. The pointer should contain a set of factors (one for each marker) where each factor contains the labels for the alleles separated by the ‘/’ character.
Marker names
A text defining the marker names for each set of genotype scores.
Linkage groups
A factor defining the different linkage groups (or chromosomes).
Positions within linkage groups
A variate specifying the positions of each marker within the linkage groups.
Marker sets
A factor defining marker sets, to specify different types of marker.
Id labels for genotypes
A text specifying labels for the genotypes that correspond to the labels for the genotypes loaded with the phenotypic means. This data structure can be used to enable correct matching for the genotypes across the genotypic and phenotypic data.
Parental information
A pointer specifying the parental information for the marker genotype scores.
Labels for parents
A text specifying labels for the parents that correspond to the text structures in the parental information pointer.
Subpopulation groups
A factor specifying subpopulation groups for association analysis.
Kinship matrix
A symmetric matrix of kinship information for association analysis.
Action Icons
| Clear | Clear all fields and list boxes. | |
| Help | Open the Help topic for this dialog. |
See also
- QTL data space for using data in QTL menus
- Adding phenotypic plot data, phenotypic means or genetic predictor data to a QTL data space.